4.4: Physical Lipid Protein Interactions
- Page ID
- 14600
Biological membranes play a prominent role in cell biology. The lipid-bilayer is composed of a large variety of lipid molecules that naturally form physical barriers creating compartments needed for cell regulation [1]. The shape of biological membranes is dictated by its lipid composition and a vast array of membrane-proteins inserted on it. The lipid environment and membrane-proteins are subject to chemical and physical interactions. Chemical interactions include effects of residues between lipids and proteins at the molecular level [3]. Physical interactions include mechanical stimuli between lipids and proteins.
In 1972, Singer and Nicolson proposed the fluid mosaic model which describes the lipids as a neutral environment to the activities of proteins [1,2]. However, recent studies suggest that lipids are strongly coupled to membrane-proteins and at times the lipid bilayer has the leading role determining the function of membrane-proteins [3, 8 ,10]. One specific case of study highlighting the physical lipid-protein interactions concerns the mechanosensitive (MS) ion channels.
Mechanosensitive Ion Channels
A membrane protein is considered mechanosensitive if its activity is controlled by membrane tension of approximately 20 nN/m. Mechanical forces can stretch, compress and bend the lipid bilayer causing membrane tension [4,5]. MS ion channels then act like mechanical force sensors transducing mechanical stimuli into important intracellular signals. In general, MS channels can be divided into two groups: prokaryotic (bacterial and archaeal) and eukaryotic (fungal, plants and animal and human) [11]. In both of these groups the function of MS channels is different. For example, in animals and human they are important in touch sensation, balance and hearing. In bacteria, MS ion channels are associated with osmoregulation control] [7,8]. MS ion channels highlight the physical lipid-protein interactions and its function depend upon mechanical membrane parameters such as tension and thickness [5]. Tension and thickness due to lipids induced perturbations to the membrane shape and the free-energy cost of such perturbations can be estimated quantitively by simple mathematical models.
Prokaryotic Mechanosensitive Ion Channels
Based on the homology of their primary sequence, MS channels in prokaryotic cells MS channels are classified into two categories: Large conductance MS channels (MscL) and small conductance MS channels (MscS). In principle MscL and MscS channels undergo the same conformation from a closed state to an open state and its thermodynamics can be described by a two-state model [9]. MS channels from MscL bacteria undergoes a large in-plane area expansion when it opens. In the case of MscL, pore gating serves as a “pressure release valve” and the in-plane expansion produces a decrease in the free energy of the membrane-channel system if the membrane is under tension and as a result it favors the open state [6,9]. To understand the free-energy change, the tension here is analogous to a gravitational field that tends to lower the weights [6]. Figure \(\PageIndex{1}\)A shows that when the channel is closed, tension is small and the weights are not moving, if the weights are lowered this produces tension and channel will open (Figure \(\PageIndex{1}\)B).

The free energy difference of the open-close channel is given as follows:
\[ ΔG = ΔG (γ=0) − γΔA \label{Equation 1}\]
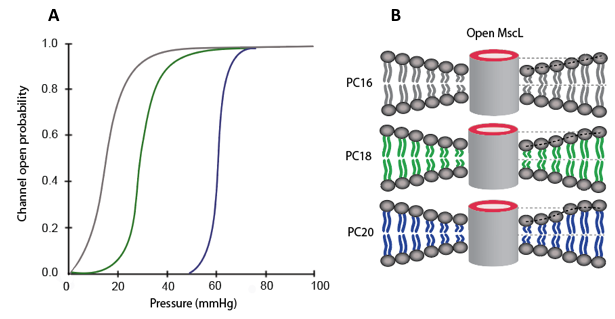
where ΔG (γ=0) is the change in free-energy at zero tension, γ is the membrane tension and ΔA is the in-plane cross sectional area. In MscL the in-plane area change ΔA has been calculated experimentally is approximately 20 nm2. Thus, the energy required to fully open the MscL channel is about 20 kT. In addition, the experimental results from MscL in Escherichia coli not only show the effects of tension but also show that lipid tail length, saturation and head group affect the channel gating properties [2,7]. The data in Figure \(\PageIndex{2}\)A shows the open probability as a function of pressure for MscL. Different traces correspond to different lengths in lipid tails. The tails have 16, 18 and 20 carbon atoms in their backbone [2,7]. Figure \(\PageIndex{2}\)B shows how the thickness of the lipid bilayer is affected by the length of lipid tails, this difference in thickness is often referred as a hydrophobic mistmatch.
Techniques used to understand the mechanism for gating of MS ion channel in prokaryotic cells include disulfide trapping, spectroscopic probing, mutagenesis screening and chemical modification [9]. In addition, patch clamp experiments have demonstrated that bacteria MscL channels gates respond to an increase in trans bilayer pressure [7].
Eukaryotic Mechanosensitive Ion Channels
In eukaryotic cells, there are four types of MS channels which include members of DEG/EnaC, 2P-type K, TRP and Piezo channel families [11]. In particular, Piezo 1 ion channels similar to MscL and MscS channels to open only requires the effects of lipid bilayer alone without the requirement of other cellular components [11].
Piezo 1 ion channels are sensitive to mechanical forces and transduce them into electrical signals in sensory neurons to achieve control in touch sensation, balance and cardiovascular regulation. The essential biophysical characteristics of Piezo 1 ion channels include control gating through mechanosensitive forces and cation selectivity [6]. It has been suggested that Piezo 1 channels sense force directly through the lipid membrane in a similar way MS channel from MscL bacteria. However, Piezo 1 channels have narrow pores in comparison to MscL channels and the effects on the membrane are not directly associated with in-plane area expansion, instead Piezo channels deform the membrane by producing local curvature (dome-shape) [6]. The expansion of the membrane-channel system occurs when the dome shape becomes flatter thus, the free energy difference between open-close channel is given as follows:
\[ ΔG = (ΔG_{prot} + ΔG_{bend} ) − γΔA_{proj} \label{Eq.2 }\]
where \(ΔG_{prot}\) and \(ΔG_{bend}\) refer to free energy differences intrinsic to protein gating and membrane bending, γ is the membrane tension and \(ΔA_{proj}\) the change in area of the dome that is projected onto the membrane plane (Figure \(\PageIndex{3}\)B). Due to its complex structure, the values of A have not been determine experimentally. For instance, if the Piezo 1 ion channel becomes completely planar, the projected area \(ΔA_{proj}\) is about 20 nm2 [6].Then the energy required to open this channel is approximately 40 kT. However, if Piezo 1 channels only flattens partially then the \(ΔA_{proj}\) should be less than 120 nm2 and the energy will be less than 40 kT. In addition, if ΔG > 20 kT will imply that Piezo 1 channels are more sensitive to tension than MscL.

Techniques used to study MS ion channels in eukaryotic cells include crystallography, cryo-EM, SDSL EPR, SDFL FRET spectroscopy, path fluorometry, mass spectroscopy and computer modelling [11].
Summary
- In prokaryote and eukaryotic cells, MS channels senses stimuli from membrane tension. Specifically, MscL, MscS and Piezo 1 channel requires only the effect of the lipid bilayer without the requirement of other cellular components.
- In the case of MscL the open channel mechanosensitive is due to couple pore opening to in-plane area expansion. However, in the Piezo channel, flattening of the channel and its nearby surrounding membrane causes membrane plane expansion by transferring out-of-plane membrane area into the membrane plane.
- Simple mathematical models can be used to understand and get insights of how the perturbations to the membrane shape affect the function of mechanosensitive ion channel.
References
- D.M. Engelman, Membranes Are More Mosaic Than Fluid, Nature (London) 438, 578 (2005).
- Phillips et al., 2008 Phillips RKJ, Theriot J, Garcia HG. Physical Biology of the Cell. Garland Sci. 2013.
- Phillips et al., 2009R. Phillips, T. Ursell, P. Wiggins, P. Sens. Emerging roles for lipids in shaping membrane-protein function.Nature, 459 (2009), pp. 379-385
- Brown, M. F. Soft Matter in Lipid–Protein Interactions. In Annual Review of Biophysics; Dill, K. A., Ed.; Annual Reviews: Palo Alto, 2017; Vol. 46, pp 379– 410
- Reeves, D., Ursell, T., Sens, P., Kondev, J. & Phillips, R. Membrane mechanics as a probe of ion-channel gating mechanisms. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 78, 041901 (2008).
- Y.R. Guo, R. MacKinnon. Structure-based membrane dome mechanism for Piezo mechanosensitivity Elife, 6 (2017), 10.7554/eLife.33660
- Perozo,E.,Kloda,A.,Cortes,D.M.&Martina,B.Physicalprinciplesunderlyingthe transduction of bilayer deformation forces during mechanosensitive channel gating. Nature Struct. Biol. 9, 636–637 (2002).
- Brohawn, S. G., Su, Z. & MacKinnon, R. Mechanosensitivity is mediated directly by the lipid membrane in TRAAK and TREK1 K+ channels. Proc. Natl Acad. Sci. USA 111, 3614–3619 (2014).
- Zhang XC, Liu Z, Li J. From membrane tension to channel gating: a principal energy transfer mechanism for mechanosensitive channels. Protein Sci 25:1954 –1964 (2016).
- Pivetti, C. D., Yen, M.-R., Mille, S., Busch, W., Tseng, Y.-H., Booth, I. R. and Saier, M. H., Jr (2003). Two families of mechanosensitive channel proteins. Microbiol. Mol. Biol. Rev. 67, 66-85.
- Battle, A.; Ridone, P.; Bavi, N.; Nakayama, Y.; Nikolaev, Y.A.; Martinac, B. Lipid–protein interactions: Lessons learned from stress. Biochim. Biophys. Acta (BBA)-Biomembranes 2015, 1848, 1744–1756.

