5.3: Styrene Maleic Acid Lipid Particles (SMALP) Technology
- Page ID
- 1351
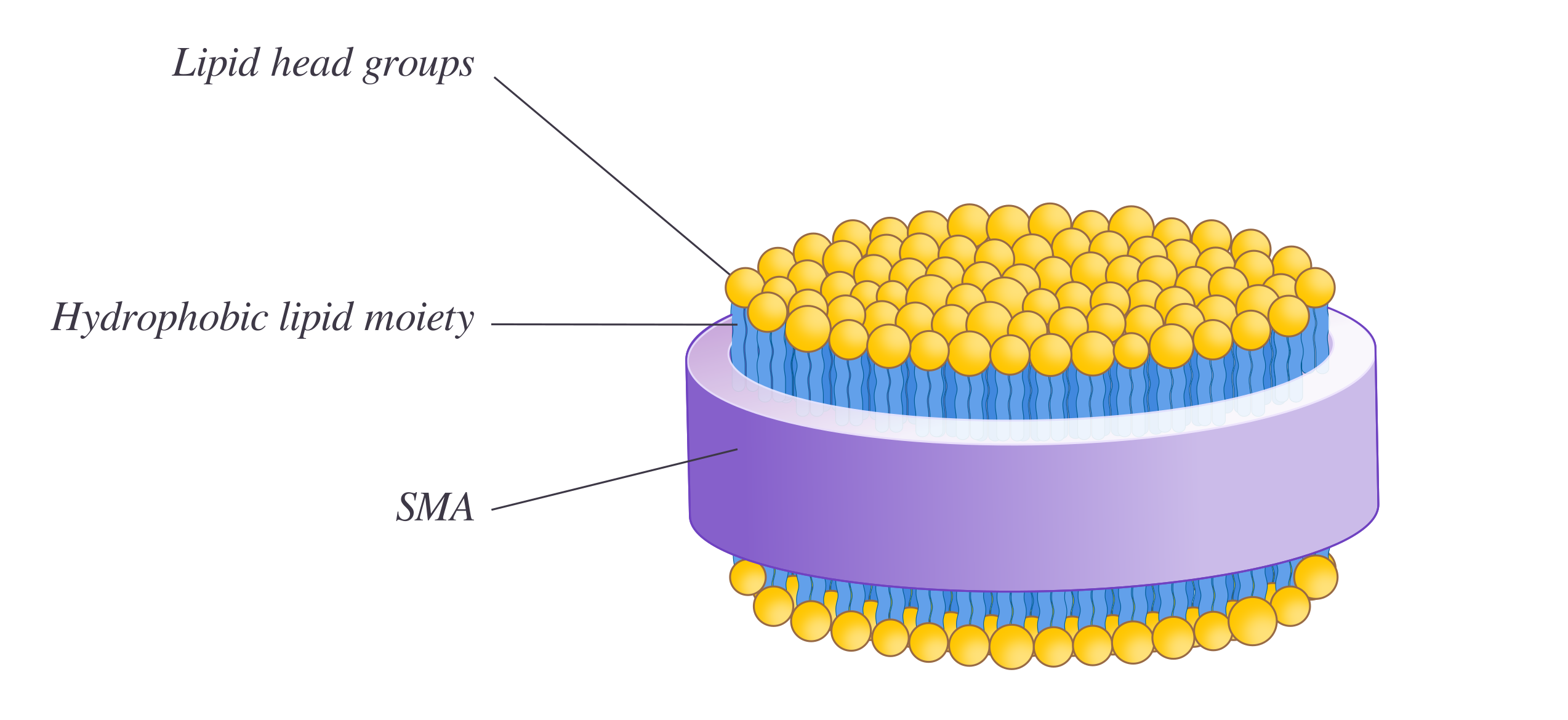
Styrene maleic acid lipid particles (SMALPs) are disc-shaped lipid assemblies on the nano-scale made from the interaction of membrane bilayers and styrene maleic acid (SMA) copolymer. SMALP technology holds great potential as a biochemical tool as it allows solubilization of membrane bilayers and its embedded constituent into discs without any use of detergents. SMALPs are thought to be superior to other disc-forming techniques eg. nanodisc as they can form directly from the native biological membrane bilayer and preserve the lipids surrounding the protein of interest. Disc-structure can potentially improve the understanding of membrane proteins by enabling biophysical studies eg. short-angle scattering, dynamic light scattering. Since the use of detergents often destabilizes proteins, SMALPs are advantageous to other established membrane systems.
Styrene Maleic Acid (SMA)
SMA copolymer is formed from polymerization of a mixture of styrene and maleic anhydride in various ratios (3:1 and 2:1 being the most common). The anhydride moieties can subsequently be hydrolyzed to maleic acid (Figure \(\PageIndex{2}\)). The alternating hydrophobic residues (styrene) and hydrophilic (maleic acid) is thought to be determining for the SMA properties of membrane solubilization.
SMA can be purchased from a range of vendors hereunder Cray Valley, Polyscope Polymers (Xiran) and Sigma-Aldrich (3:1 SMA, Lipodisq).
SMA lipid particles (SMALPs)
After originally being patented as Lipodisq, a drug delivery tool for hydrophobic pharmaceuticals1, SMALPs with integrated membrane proteins were described by Knowles and coworkers in 20092. The group showed SMALP integration of both the alpha-helical (7TM) protein bacteriorhodopsin (bR) and the beta-barrel protein PagB from dimyristoyl PC liposomes. After purification via Ni2+-affinity chromatography, structural analysis dynamic light scattering (DLS), transmission electron microscopy (TEM) and circular dichroism (CD) spectroscopy was carried out. TEM showed nanoparticles of approximately 11 nm in diameter for both proteins, which was consistent with DLS data2 and results from later studies (12 nm by3–5, 9 nm by1). Furthermore, CD spectroscopy showed that both proteins maintained structure after integration into SMALPs while the absorbance of retinal in bR slightly shifted. A similar shift is, however, also seen in detergent. The author attributed this shift to the monomeric quaternary structure in SMALPs and detergent compared to the trimeric structure in the native purple membranes2.
Table one contains an overview of the literature concerned on this page and the techniques and concepts that is touched upon.
Table 1 Overview of literature
SMALP formation
Sheidelaar and coworkers5 investigated the kinetics of SMALP formation (2:1 SMA) from large unilamelar vesicles (LUVs) by monitoring the drop in optical density when LUVs solubilize. The results showed that higher temperature, relative to the lipid Tm, and higher SMA concentration both increased kinetics of SMALP formation from di-saturated PC LUVs. This trend was consistent with different various chain lengths. Furthermore, the study showed that the size of the discs was similar regardless of chain length.5 The exact formation of SMALPs are not entirely clear. Scheidelaar and coworkers propose a three-step model where
- SMA binds the membrane surface,
- destabilize the bilayer before
- the formation of the dis.c5
Lipid saturation and lipid packing: The lipid packing of the bilayer was shown to strongly influence the SMALP formation kinetics. Even though saturated lipids showed faster kinetics at T>Tm, LUVs of mono-unsaturated lipids (that are in the fluid phase at room temperature) showed slower kinetics of SMALP formation.
Accordingly, bilayers of PC with two mono-unsaturated fatty acid chains showed slower kinetics than bilayers of PC with only one mono-unsaturated chain (and one saturated chain) whilst bilayers from PC with two di-unsaturated chain showed similar kinetics (where x is the chain length). Thus the extra double bond in the fatty acid chains does not have a large impact compared to going from saturated to unsaturated. All except di-22:1 PC bilayers, formed SMALPs within 10 min at 30°C. The importance of lipid packing was underlined by results showing a negative correlation between lateral membrane pressure and SMA-mediated solubilization via experiments with bilayers of lipid mixtures with different spatial properties.
Salt concentration: The majority of biological membrane lipids have anionic head groups. Electrostatic repulsion between these head groups and the negatively charged (at physiological pH) maleic acid moieties of SMA might occur. Monitoring SMALP formation from LUVs containing the anionic lipid di-C14:0 PG showed that formation was inhibited in these LUVs in the absence of salt. SMALPs did form in LUVs with mixed lipids. The absence of salt is, however, generally inhibitory to SMA insertion into membranes as it shields the charge repulsion5. Sodium chloride and pH buffer (Tris-HCl pH 8) is the only other necessary components to form SMALPs except for SMA and a membrane bilayer eg. a vesicle.
SMALP formation from cell membranes
Sheidelaar and coworkers5 demonstrated SMALP formation from Escherichia coli membrane fragments to be achievable with excess SMA. Furthermore, the group showed that the membranes fragments, the SMALPs and fatty acids/lipids in the aqueous phase all contained similar relative amounts of PG, PC and cholesterol, thus indicating that SMALPs are not selective for specific lipid types in the membrane bilayer.
Swainsbury and coworkers4 successfully purified the photoreaction center from the bacterium Rhodobacter sphaeroides directly by SMALP formation from the bacterial membrane with subsequent Ni2+-affinity chromatography. The study showed that it was possible to purify membrane proteins/complexes in an active state, in its immediate native lipid environment and underlined that it could be used to study the biochemistry and biophysics in the SMA scaffold. This is interesting as potentially can enable elucidation of the role of nearby lipids for protein function as eg. non-annular lipids.
Conclusion
Styrene maleic acid lipid particles efficiently form disc-shaped nanostructures from intact lipid bilayers from both vesicles and cells with only minor dependency on salt concentration. Biophysical studies can be carried out on proteins embedded in SMALPs in order to study these proteins in the native lipid environment. SMALPs can potentially strengthen the understanding of membrane lipids that are essential for membrane protein function.
References
- Orwick, M. C. et al. Detergent-free formation and physicochemical characterization of nanosized lipid-polymer complexes: Lipodisq. Angew. Chemie - Int. Ed. 51, 4653–4657 (2012).
- Knowles, T. J. et al. Membrane proteins solubilized intact in lipid containing nanoparticles bounded by styrene maleic acid copolymer. J. Am. Chem. Soc. 131, 7484–7485 (2009).
- Orwick-Rydmark, M. et al. Detergent-free incorporation of a seven-transmembrane receptor protein into nanosized bilayer lipodisq particles for functional and biophysical studies. Nano Lett. 12, 4687–4692 (2012).
- Swainsbury, D. J. K., Scheidelaar, S., van Grondelle, R., Killian, J. A. & Jones, M. R. Bacterial Reaction Centers Purified with Styrene Maleic Acid Copolymer Retain Native Membrane Functional Properties and Display Enhanced Stability. Angew. Chemie Int. Ed. 53, 11803–11807 (2014).
- Scheidelaar, S. et al. Molecular Model for the Solubilization of Membranes into Nanodisks by Styrene Maleic Acid Copolymers. Biophys. J. 108, 279–290 (2015).

