5.1: Model Membranes vs. Biological Membranes
( \newcommand{\kernel}{\mathrm{null}\,}\)
The study of cells has been a challenging, active area of research since the seventeenth century when Robert Hooke first observed them through his compound microscope. [1] Cells consist of cytoplasm, which contains biomolecules such as nucleic acids and proteins, and is enclosed by a plasma membrane. This membrane plays an essential role in cellular protection as well as in the control and the transport of ions, nutrients and a variety of small molecules through protein channels or processes like diffusion and endocytosis.
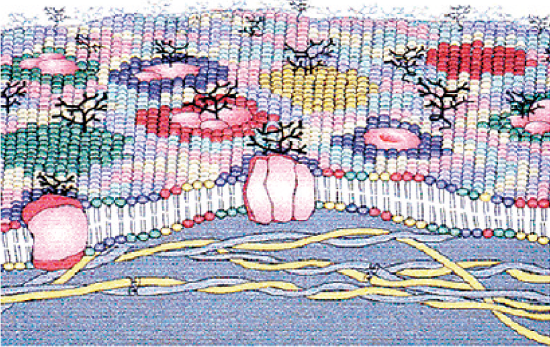
One of the key developments towards determining the structure of the plasma membrane was the proposal of the “Fluid Mosaic Model” by SJ Singer and GL Nicolson in 1972. The model states that membranes are composed of a phospholipid bilayer backbone with embedded cholesterol, proteins, and carbohydrates that gives the membrane its fluid character. [11] This composite structure allows the membrane to perform multiple functions such as molecular recognition, enzymatic catalysis, cellular adhesion and membrane fusion. The most important evolution of this model happened in 1997 with the works of Simons et al. and of Brown et al. who proposed that membrane lipids were organized into phase-separated micro-domains, called lipid rafts, with a different local composition and a molecular dynamic from the surrounding liquid crystalline phase. The validity of this hypothesis was a hotly debated topic for more than a decade but has since been demonstrated by a lot of researchers and has led to the development of technologies for detecting lateral heterogeneity in biological membranes. [10] A comprehensive understanding of all components of the plasma membrane and their synergistic workings could help improve our understanding of the evolution of life on earth and play a crucial role in biological advancement in the future.
Biological Membranes: Composition and Functions
Biological membranes have complex and varied compositions of lipids and proteins which make the determination of their exact composition in a cell quite difficult. That being said, their primary components are well established.
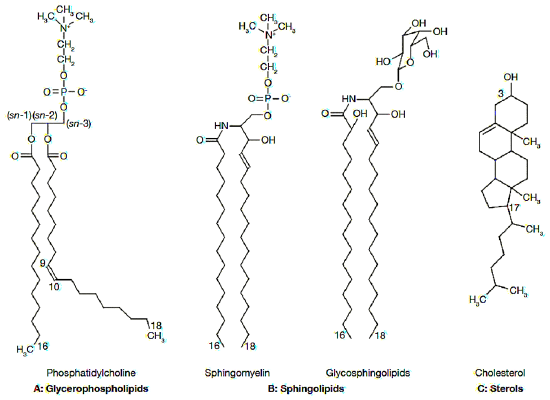
The first major components of biological membranes are lipids. All biological membranes contain lipid bilayers as their basic structural unit. Lipid bilayers are sheet-like assemblies of thousands of amphiphilic lipid molecules held together by hydrophobic interactions between their acyl chains. Membrane lipids can be divided into three groups based on their chemical structure: glycerol-based lipids (phospholipids), ceramide-based sphingolipids, and sterols. (Figure 5.1.1a) These constituents are not homogeneously arranged in the cell plasma membrane but actually present as complex lateral micro-domains. (Figure 5.1.1b) This variability in the nature of lipid arrangement, in addition to significant differences in physical properties like cross-sectional area, unsaturation, fluidity, electric charge, molecular weight, is the reason that lipid membranes are extremely complex structures. Their association with other membrane components like proteins and carbohydrates adds to the difficulty of studying the membrane’s structure. [12]
The second major components of cell membranes are the proteins. They constitute a large proportion of a typical cell membrane and perform a variety of functions [1]. Membrane proteins can associate with membranes in a variety of ways such as having domains that traverse the membrane, domains that associate with only one face of the membrane, chemical groups that anchor them to the membrane, or domains that associate with other membrane-bound proteins. The low permeability of cell membranes to everything except water, small uncharged molecules, and some hydrophobic signalling molecules requires that they possess machinery to facilitate the exchange of molecules between the cytoplasm and their external environment. Membrane proteins facilitate these types of phenomena. For example, proteins that act as channels to allow the rapid transport of ions and other molecules are quite common [2]. Other types of membrane proteins act as receptors to perceive external or internal signals, docks for recognition and fusion of vesicles carrying cellular cargo, enzymes that catalyze specific chemical reactions at membranes, channels which facilitate the transport of other proteins across a membrane, and can associate in protein complexes of varying size and composition [1]. Clearly, there is a vast amount of diversity in cell membranes in terms of lipid and protein composition and how these components influence the properties of the membrane.
Carbohydrates covalently linked to proteins (glycoproteins) or lipids (glycolipids) are also a part of cell membranes, and function as adhesion and address loci for cells thereby aiding in cell-cell contact, recognition of self (via presentation of cell-specific carbohydrates that are recognized and ignored by immune cells), and communication. [17]
Cell membranes are highly dynamic structures. They are the interface between a cell and its environment. They perceive signals from the surroundings, transduce the signals, and convert them into cellular responses. They are like busy highways in cells in the sense that information is being rapidly being transmitted through and across them. Living cells also constitutively modify their membranes as a part of normal cell function and in specific manners in response to external stimuli. Another very important property of cell membranes is that they possess electrical and chemical gradients produced by resident proteins. These gradients are essential for functions of living cells such as energy production, solute uptake, and signal transduction.
Model Membranes Systems
Model organisms are often used by scientists to help understand biological processes because they are easy to breed, maintain and manipulate. Similarly, model membranes can be used to gain insight into the properties and functions of different components of biological membranes [3]. The complexity of cell membranes, and the difficulty of isolating them and maintaining their native physiological conditions makes it challenging to extract meaningful data from experiments on whole cells. Thus, scientists frequently use artificial membranes which have greatly simplified structures and composition to make experiments more tractable.
Model membranes are a useful tool to probe the behavior of proteins and lipids in a membrane by isolating different aspects of membrane function and studying them in detail to get some insight about the bigger picture. They can be used to study things like the structure and functions of type of lipids, effect of curvature membrane protein complexes, ion channels, and also study the interactions of lipids with drugs or other nanoparticles [3]. The most well-known and common biomimetic systems used for such purposes are lipid monolayers, lipid vesicles and supported lipid bilayers. While each of these systems exhibits advantages and disadvantages, they all mimic the lipid arrangement of natural cell membranes.
Lipid Monolayers
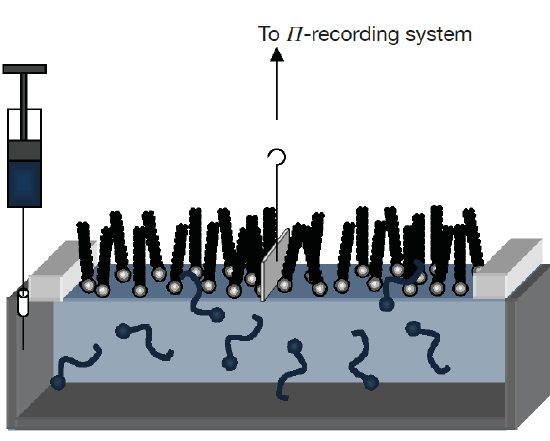
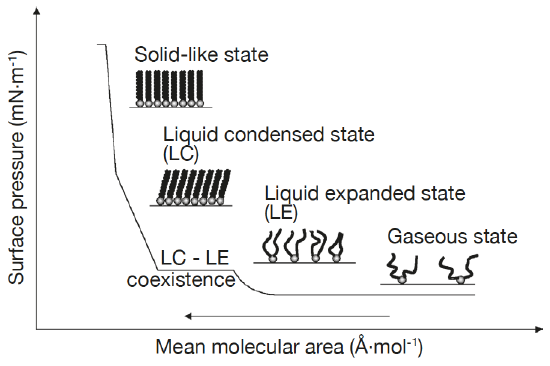
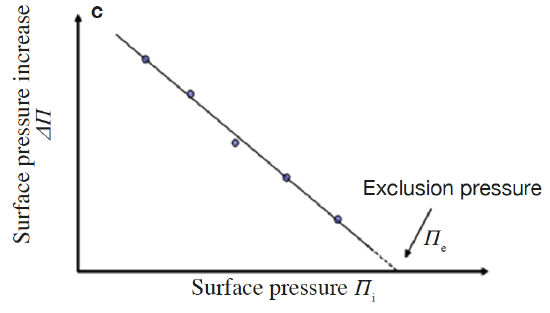
Lipid monolayers which are half of a bilayer, are formed by spreading amphipathic molecules on the surface of a liquid. They provide a simple model for evaluating membrane insertion of compounds and for studying lipid-lipid interactions by changing parameters such as the nature and the packing of the spread molecules, the composition of the subphase (pH, ionic strength) and temperature. Compression isotherms are often used to get information about lipids phase (gel or liquid) and to study the nature of lipid mixtures for phenomena like phase separation. These are obtained by measuring the surface pressure (Π) of the interfacial film as a function of the mean molecular area (A) in a Langmuir-Blodgett trough. (Figure 5.1.3a,b) To study the insertion of molecules such as drugs into a membrane, a monolayer is held at an initial pressure using the barriers of the Langmuir trough and the change is pressure on insertion of solubilized compound of interest is noted after the system pressure stabilizes. By plotting the surface pressure increase (ΔΠ) observed as a function of the initial surface pressure of the lipid monolayer, we get a maximum insertion pressure (Πm) which reflects the penetration power of the molecule of interest into a well-defined 2-D model membrane. (Figure 5.1.3c) [10]
Lipid Vesicles
Lipid vesicles or liposomes are the simplest models of closed cell membranes. They are spherical lipid bilayers with an internal aqueous compartment and can be formed in different sizes, SUVs (small unilamellar vesicles, 20-50 nm in diameter), LUVs (large unilamellar vesicles, 100-500 nm in diameter), and GUVs (giant unilamellar vesicles, 10-100 µm in diameter). [12,13] GUVs are closest in size to actual cells and are usually created using well-defined mixtures of pure lipids but it is unclear how the behavior of these model systems compares to actual biological membranes which are much more complex mixtures of lipids, proteins, and saccharides. (Picture 4a) Giant plasma membrane vesicles (GPMV’s), or blebs, are a more direct source of biological lipid material. In response to chemical stress, the plasma membrane of cells protrudes and detaches to form these blebs, which are similar in size to the GUVs. The lipid composition of blebs is similar to that of the parent cell such that they also contain membrane proteins which diffuse into the detached area during their formation. [14]
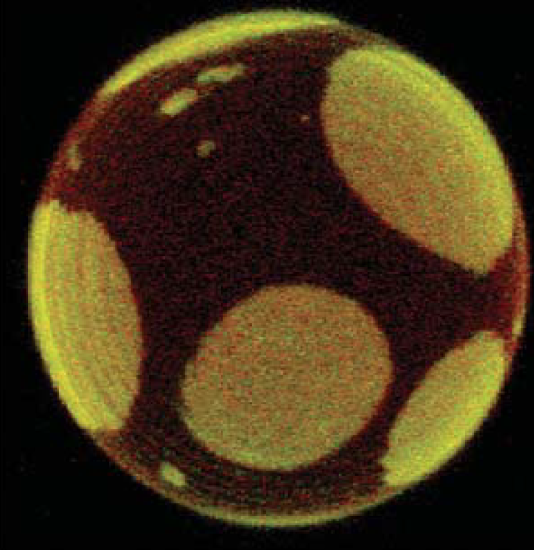
The most common applications of GUVs and GPMVs are to study phase behavior, (Figure 5.1.4a) effects of membrane permeability barriers and curvature. [13] They are also used to investigate membrane processes such as membrane fusion, molecular recognition, cell adhesion, and membrane trafficking. The size of these giant vesicles allows the use of optical microscopy techniques such as fluorescence or confocal microscopy, as well as the micromanipulation of individual vesicles. Although these techniques have a lower lateral resolution than AFM, they allow the investigation of molecular interactions with lipid vesicles in a bulk solution whereas AFM requires the fusion of lipid vesicles onto a solid support. [10]
One drawback of using vesicles is that they are metastable structures offering poor long-term stability. Upon aging, vesicle dispersion may aggregate, fuse or evolve to the thermodynamically stable two-phase region (consisting of a lamellar phase dispersed in large excess of solvent) from which they were formed. However, depending on the composition and the size of lipid vesicles as well as on the environmental parameters (temperature, pH, ionic strength, presence of external molecules and ions), these thermodynamically non-stable systems can be stable for prolonged periods of time (up to several months) and are then suitable model membranes for investigating membrane properties and biological processes.[10]
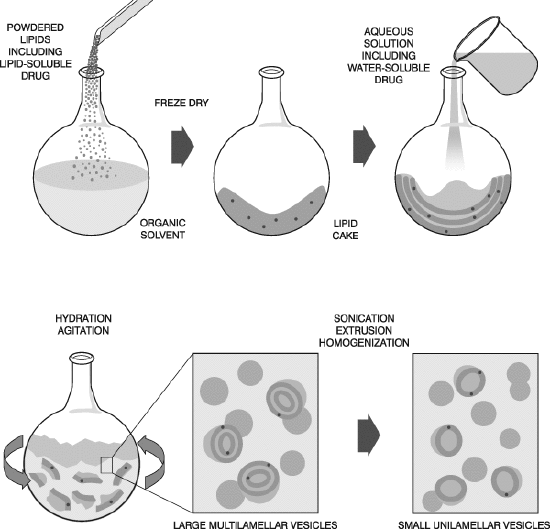
SUVs and LUVs are prepared by dissolving lipids in organic solvents. Then the solvents are evaporated under a vacuum to form thin films of lipids at the bottom of a round bottom flask. The dried lipid films are resuspended in a desired buffer to form liposomes. The liposome suspension is subjected to vortexing until complete dissolution of the lipid film, followed by ultrasonication. This procedure usually results in multi-lammelar liposomes (MLVs), which are subjected to 3–5 cycles of freeze/ thawing for the uniformity of liposome size. GUVs are prepared by electroformation. In this method, the lipid film is dried under an oscillating electric field. Typically, a standard wave generator is used to apply 1 V at 10 Hz between electrodes onto which a thin film of lipids has been dried in the presence of water to form GUVs. [12] (Picture 4b)
Tethered Vesicles
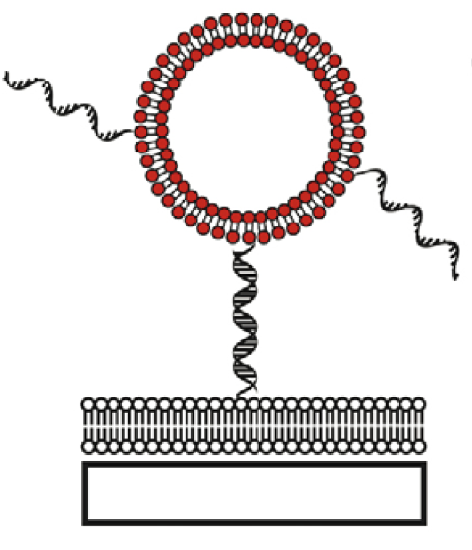
Several groups of researchers have tethered lipid vesicles to solid supported bilayers using DNA hybridization or biotin–streptavidin recognition elements. By doing so the proteins incorporated into tethered vesicles are shielded from the solid support by the presence of a supported bilayer (SLB). DNA-tethered vesicles are constrained to diffuse in two dimensions in a plane parallel to the surface. Interactions between vesicles with reactive membrane components like DNA or proteins can be observed as vesicles diffuse and collide with each other. Vesicles tethered with biotin–streptavidin coupling via biotinylated lipids are often observed not to diffuse, so this system is not useful for studying interactions between tethered vesicles but this can be an excellent system for encapsulating reactants in small volumes for fluorescence measurements and to look at reactions of these tethered vesicles with free vesicles flowed over the surface from bulk solution. [14] (Picture 4c)
Lipid Bilayer – Supported and Tethered
Supported lipid bilayers (SLBs) are stable model membranes consisting of a flat lipid bilayer supported on a solid surface such as mica, glass or silicon oxide wafers, with the polar head facing the support. Unlike vesicles, these model membranes are easy to prepare and control the overall composition and the lipid asymmetry. [10] SLBs are used to predict phase behavior and the molecular organization of biological membranes. They have also been used for investigating the molecular interactions of drugs with cell membranes [10]. Changes in structure, morphology, and surface chemistry of SLBs following interaction with drugs or drug delivery systems can be investigated using various techniques, such as X-ray scattering, scanning electron microscopy, atomic force microscopy (AFM), transmission electron microscopy, Fourier transform infrared resonance (FTIR), and X-ray photoelectron spectroscopy (XPS) [12].
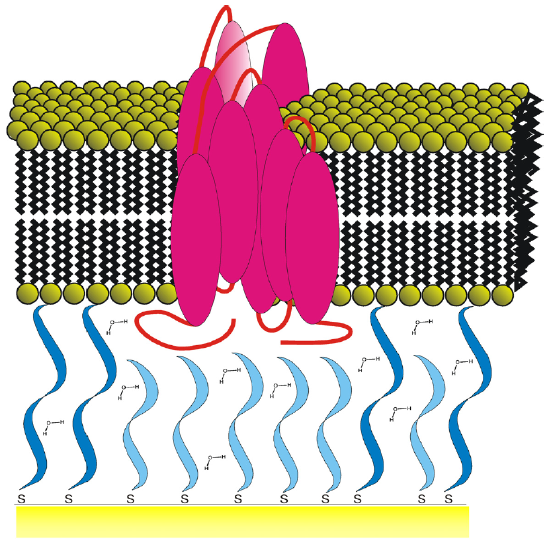
One of the main drawbacks of using classical supported lipid bilayers is the proximity between the lipid bilayer and the solid substrate. This may affect the membrane properties like the mobility of membrane components or the incorporation of transmembrane proteins. One way to counter this problem is by assembling bilayers on softer supports such as polymer cushions. [10] Another way is to use tethered bilayer lipid membranes (t-BLMs) made up of a lipid bilayer spaced from the solid surface using spacer molecules (Figure 5c) .The spacer allows the bilayer to retain the dynamic properties like lateral diffusion of the lipid molecules in the bilayer. Studies of the diffusion coefficient of individual lipids within the bilayer by fluorescence recovery after photo-bleaching (FRAP) measurements have demonstrated the high lateral mobility of these lipids in the tethered system. The spacer layer also provides space necessary to prevent undesirable interactions between membrane proteins and the substrate, like denaturation. In addition to these aspects, the tether also gives this system the mechanical and chemical stability and robustness that is needed for biosensor applications. [15]
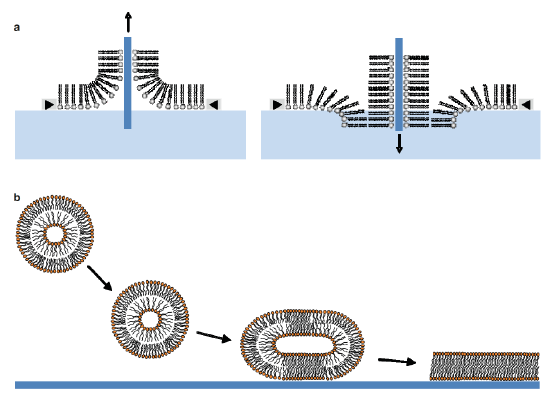
Different techniques are commonly used to prepare SLBs. The first one is the LB technique. After the transfer of a lipid monolayer spread at the air-water interface of a Langmuir trough onto a solid support, the same support is immersed a second time through the interface in order to obtain a supported lipid bilayer. A second method for preparing SLBs is the fusion of lipid vesicles onto a solid support. This is done by heating a SUV suspension in contact with the support at temperatures above the lipid phase transition. The fusion process involves the adsorption of the lipid vesicles on the surface, followed by their deformation, their flattening and their rupture. The fusion of the edges of the bilayer patches through hydrophobic interactions gives rise in final to a continuous supported lipid bilayer. [10] (Figure 5.1.5a,b)
Limitations of Model Membrane-Based Studies
One of the biggest questions to be aware of when studying model systems is, “Do the findings obtained from these experiments have bearing on what actually occurs in a living system?” As described in the first two sections, biological membranes are highly complex structures with a great deal of organization. Over the last century, model membranes have been proved to play a considerable role in the elucidation of the structure and the properties of biological membranes. While the simplification of the membrane system is crucial for the analysis of specific molecular interactions at the membrane level, it can also be an obstacle to the accurate understanding of some membrane functions. Most membrane models can only involve up to 3-4 lipid species when actual plasma membranes have more than thousands. The preparation techniques for making vesicles is quite difficult and cannot mimic the asymmetry of lipids in the two bilayer leaflets like in real membranes which is crucial to many membrane functions.
Another limitation of these model systems lies on the fact that it is quite challenging to reconstitute proteins into model membranes. Model membranes also do not contain cytoskeletal components which strongly participate to the lipid and protein diffusion across the cell surface and consequently to the phase behavior of cell membranes. [10]
Concluding Remarks
Despite the limitations of model-based studies, they have proven to be very useful in understanding basic biophysical properties of biological membranes. They provide a foundation upon which new hypotheses can be generated and tested in living cells. A solid foundational understanding of simple synthetic membranes should allow scientists to develop experimental and computational tools to test more complex properties of biological membranes.
References
- Alberts, J., Lewis, Raff, Roberts, Walter, Molecular Biology of the Cell. 2008. 5.
- Campbell, R., Urry, Cain, Wasserman, Minorsky, Jackson, Biology. 2008. 8.
- Szoka Jr, Frank, and Demetrios Papahadjopoulos. "Comparative properties and methods of preparation of lipid vesicles (liposomes)." Annual review of biophysics and bioengineering 9.1 (1980): 467-508.
- Van Meer, Gerrit, Dennis R. Voelker, and Gerald W. Feigenson. "Membrane lipids: where they are and how they behave." Nature reviews. Molecular cell biology 9.2 (2008): 112.
- Rigaud, Jean-Louis, Bruno Pitard, and Daniel Levy. "Reconstitution of membrane proteins into liposomes: application to energy-transducing membrane proteins." Biochimica et Biophysica Acta (BBA)-Bioenergetics 1231.3 (1995): 223-246.
- Szoka, Francis, and Demetrios Papahadjopoulos. "Procedure for preparation of liposomes with large internal aqueous space and high capture by reverse-phase evaporation." Proceedings of the National Academy of Sciences 75.9 (1978): 4194-4198.
- Olson, F., et al. "Preparation of liposomes of defined size distribution by extrusion through polycarbonate membranes." Biochimica et Biophysica Acta (BBA)-Biomembranes 557.1 (1979): 9-23.
- Kikuchi, Shingo, et al. "Uncovering the protein translocon at the chloroplast inner envelope membrane." Science 339.6119 (2013): 571-574.
- Schütz, Gerhard J., Hansgeorg Schindler, and Thomas Schmidt. "Single-molecule microscopy on model membranes reveals anomalous diffusion." Biophysical journal 73.2 (1997): 1073-1080.
- Eeman, Marc, and Magali Deleu. "From biological membranes to biomimetic model membranes." Biotechnologie, Agronomie, Société et Environnement 14.4 (2010): 719.
- Boundless. "Fluid Mosaic Model." Boundless Biology Boundless, 26 May. 2016. Retrieved 19 May. 2017 from www.boundless.com/biology/te...del-327-11464/
- Peetla, Chiranjeevi, Andrew Stine, and Vinod Labhasetwar. "Biophysical interactions with model lipid membranes: applications in drug discovery and drug delivery." Molecular pharmaceutics 6.5 (2009): 1264-1276.
- Mouritsen, Ole G. "Model answers to lipid membrane questions." Cold Spring Harbor perspectives in biology 3.9 (2011): a004622.
- Chan, Yee-Hung M., and Steven G. Boxer. "Model membrane systems and their applications." Current opinion in chemical biology 11.6 (2007): 581-587.
- Knoll, Wolfgang, et al. "Tethered bimolecular lipid membranes—A novel model membrane platform." Electrochimica Acta 53.23 (2008): 6680-6689.
- https://avantilipids.com/tech-suppor...e-preparation/
- Alphonsus, C. S., and R. N. Rodseth. "The endothelial glycocalyx: a review of the vascular barrier." Anaesthesia 69.7 (2014): 777-784.
Contributors and Attributions
- Lucas McKinnon, Plant Sciences, UC Davis
- Deepshika Gilbile, Chemical Engineering, UC Davis